Protein Engineering Tools: Shaping the Future of Life Science
Proteins are the building blocks of life, driving countless biological processes. But what if we could redesign them to perform even better? Welcome to the world of protein engineering, where biotechnologists use cutting-edge tools to create proteins with enhanced functions. From life-saving drugs to eco-friendly industrial enzymes, protein engineering is revolutionizing life sciences and associated technology.
In this blog post, we’ll explore the tools and techniques powering protein engineering. We’ll explore rational design, semi-rational design, and random approaches. In addition, how they’re being used to solve real-world problems. Whether you’re a biotechnologist, life sciences student, or curious reader, this post will give you a front-row seat to the future of protein engineering (Figure 1).
What is Protein Engineering?
The first or basic question which come in our mind about proteins engineering is “What is protein engineering?” Let see some basic about protein engineering. Protein engineering is the science of designing and modifying proteins to achieve specific goals. By altering amino acid sequences, biotechnologists can create proteins with improved stability, activity, or entirely new functions. This field has applications in medicine, agriculture, energy, and more.
There are three main approaches to protein engineering:
- Rational Design: Precise, knowledge-driven modifications.
- Semi-Rational Design: A blend of targeted and exploratory methods.
- Random Approaches: Untargeted exploration of protein sequence space.
Each approach has unique tools and techniques, which we’ll explore in detail.
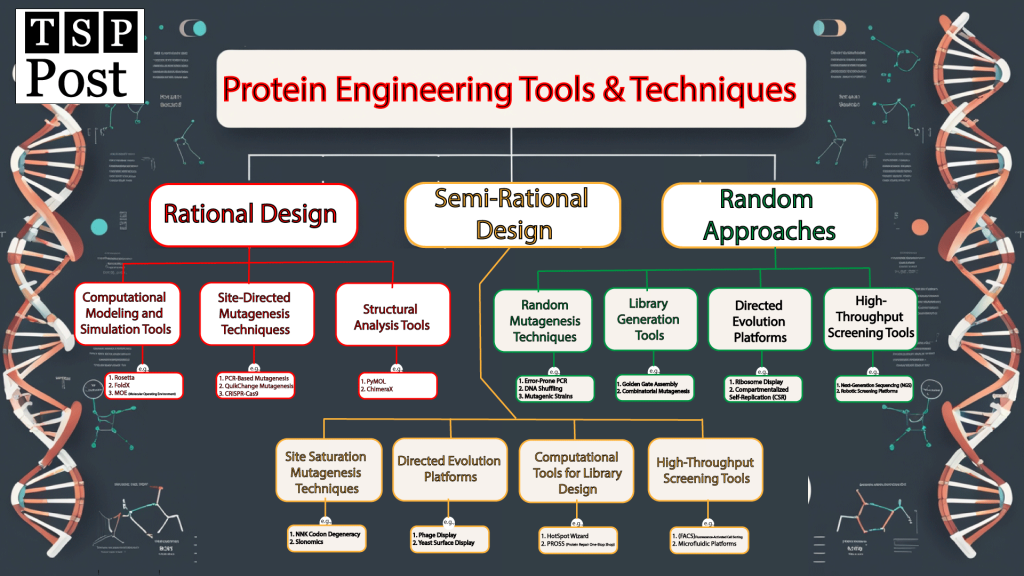
Rational Design: Precision at Its Best
Rational design is like a surgeon’s scalpel—precise and targeted. It relies on detailed knowledge of a protein’s structure and function to make specific changes. Let’s look at the tools that make this possible (Figure 1).
Computational Modeling and Simulation Tools
Computational tools are the backbone of rational design. They help biotechnologists to predict how changes will affect a protein’s behavior. The following tools are used for this purpose.
- Rosetta: This suite of algorithms predicts protein structures and identifies optimal amino acid changes. It’s like a GPS for protein engineering.
- FoldX: FoldX predicts how mutations impact protein stability and interactions. It’s a go-to tool for designing stable proteins.
- MOE (Molecular Operating Environment): MOE is used for molecular modeling and docking. It helps researchers to understand protein-ligand interactions.
Site-Directed Mutagenesis Techniques
Once the target is identified, biotechnologists use these techniques to introduce specific mutations.
- PCR-Based Mutagenesis: Custom primers introduce mutations during DNA amplification.
- QuikChange Mutagenesis: A popular method for creating point mutations in plasmids.
- CRISPR-Cas9: This gene-editing tool enables precise changes in protein-coding sequences.
Structural Analysis Tools
Visualizing protein structures is key to rational design.
- PyMOL: PyMOL creates stunning 3D models of proteins, helping biotechnologists to identify key residues.
- ChimeraX: This tool analyzes and manipulates protein structures, guiding the design process.
Example Applications
- Engineering T4 lysozyme for improved thermal stability using Rosetta.
- Designing insulin analogs with enhanced activity using FoldX and site-directed mutagenesis.
Semi-Rational Design: The Best of Both Worlds
Semi-rational design combines the precision of rational design with the diversity of random approaches. It’s ideal when you have partial knowledge of a protein’s structure and function (Figure 1).
Site Saturation Mutagenesis Techniques
These techniques introduce all possible amino acids at a specific site.
- NNK Codon Degeneracy: Degenerate primers introduce all 20 amino acids at a target site.
- Slonomics: A high-throughput method for generating and screening mutant libraries.
Directed Evolution Platforms
Directed evolution mimics natural selection in the lab.
- Phage Display: Protein variants are displayed on phage surfaces for screening.
- Yeast Surface Display: Yeast cells express protein variants, enabling high-throughput screening.
Computational Tools for Library Design
These tools help identify key residues for mutagenesis.
- HotSpot Wizard: Identifies residues for mutagenesis based on structural and evolutionary data.
- PROSS (Protein Repair One-Stop Shop): Predicts stabilizing mutations for semi-rational design.
High-Throughput Screening Tools
Screening large libraries is essential for semi-rational design.
- Fluorescence-Activated Cell Sorting (FACS): Screens protein variants for desired traits.
- Microfluidic Platforms: Enables ultra-high-throughput screening of enzyme activity.
Example Applications
- Engineering subtilisin for improved detergent compatibility using NNK-based site saturation mutagenesis.
- Optimizing antibody affinity using yeast surface display and FACS.
Random Approaches: Exploring the Unknown
Random approaches are like casting a wide net—they explore vast sequence space to discover novel functions. These methods are ideal when structural knowledge is limited (Figure 1).
Random Mutagenesis Techniques
These techniques introduce random mutations across the protein.
- Error-Prone PCR: Reduces polymerase fidelity to introduce random mutations during DNA amplification. For detail study visit our post “Error-Prone PCR: A Comprehensive Guide to Mutation Generation“
- DNA Shuffling: Recombines fragments of homologous genes to create chimeric proteins. For detail study visit our post “DNA Shuffling Techniques: Unlocking Genetic Innovation“
- Mutagenic Strains: Uses bacteria with error-prone DNA replication to generate random mutations.
Library Generation Tools
Creating diverse libraries is key to random approaches.
- Golden Gate Assembly: Combines DNA fragments to create diverse protein libraries.
- Combinatorial Mutagenesis: Randomizes multiple sites simultaneously to generate large libraries.
High-Throughput Screening Tools
Screening massive libraries requires advanced tools.
- Next-Generation Sequencing (NGS): Identifies beneficial mutations from large libraries.
- Robotic Screening Platforms: Automates the screening of thousands of protein variants.
Directed Evolution Platforms
These platforms link protein function to DNA replication.
- Ribosome Display: Screens protein variants in vitro without host cells.
- Compartmentalized Self-Replication (CSR): Links protein function to DNA replication for directed evolution.
Example Applications
- Evolving green fluorescent protein (GFP) variants with brighter fluorescence using error-prone PCR.
- Discovering thermostable enzymes for industrial applications using DNA shuffling and robotic screening.
Fore detail of Directed evolution see our post “Directed Evolution in Protein Engineering: Unlock New Potentials!“
Key Takeaways
- Rational design tools are precise and rely on computational modeling and targeted mutagenesis.
- Semi-rational design balances precision with diversity, using techniques like site saturation mutagenesis and directed evolution.
- Random approaches explore vast sequence space through untargeted mutagenesis and high-throughput screening.
By leveraging these tools, researchers can tailor their protein engineering strategies to achieve specific goals. Whether it’s precision, diversity, or discovery of novel functions, protein engineering is unlocking endless possibilities.
The Future of Protein Engineering Tools
As computational tools and experimental techniques advance, protein engineering is poised for even greater breakthroughs. Emerging technologies like machine learning and artificial intelligence are transforming the field, enabling faster and more accurate protein design.
From creating life-saving drugs to developing sustainable industrial processes, protein engineering is shaping the future of science. The journey has only just begun, and the possibilities are limitless.
So, the next time you hear about a groundbreaking discovery, remember: behind it all, there’s likely a team of protein engineers, meticulously designing the molecules that make it possible.
Comparative Table of Tools/Techniques and Their Applications
| Approach | Tools and Techniques | Applications | Examples |
| Rational Design | Computational Modeling: Rosetta, FoldX, MOE | Predicting protein stability, binding, and interactions. | Engineering T4 lysozymefor improved thermal stability using Rosetta. |
| Site-Directed Mutagenesis: PCR-Based Mutagenesis, QuikChange, CRISPR-Cas9 | Introducing specific mutations for targeted improvements. | Designing insulin analogswith enhanced activity using FoldX. | |
| Structural Analysis: PyMOL, ChimeraX | Visualizing and analyzing protein structures. | Identifying key residues in enzyme active sites using PyMOL. | |
| Semi-Rational | Site Saturation Mutagenesis: NNK Codon Degeneracy, Slonomics | Exploring all possible amino acid substitutions at a target site. | Engineering subtilisin for improved detergent compatibility. |
| Directed Evolution: Phage Display, Yeast Surface Display | Screening protein variants for binding or activity. | Optimizing antibody affinity using yeast surface display. | |
| Computational Tools: HotSpot Wizard, PROSS | Identifying key residues for mutagenesis. | Predicting stabilizing mutations for enzyme engineering using PROSS. | |
| High-Throughput Screening: FACS, Microfluidic Platforms | Screening large libraries of protein variants. | Identifying thermostable enzymes using microfluidic screening. | |
| Random Approach | Random Mutagenesis: Error-Prone PCR, DNA Shuffling, Mutagenic Strains | Introducing random mutations across the protein. | Evolving GFP variants with brighter fluorescence using error-prone PCR. |
| Library Generation: Golden Gate Assembly, Combinatorial Mutagenesis | Creating diverse protein libraries. | Generating chimeric proteins using DNA shuffling. | |
| High-Throughput Screening: NGS, Robotic Screening Platforms | Screening massive libraries for beneficial mutations. | Discovering novel enzymes using robotic screening. | |
| Directed Evolution: Ribosome Display, Compartmentalized Self-Replication (CSR) | Linking protein function to DNA replication for evolution. | Evolving industrial enzymes using ribosome display. |
Frequently Asked Q&A
What is protein engineering?
Protein engineering is the science of designing and modifying proteins to enhance their functions. By altering amino acid sequences, researchers create proteins with improved stability, activity, or entirely new properties for applications in medicine, industry, and biotechnology.
What are the main strategies to protein engineering?
Protein engineering utilizes three key strategies:
Random Approaches – Unbiased mutagenesis to explore vast sequence space.
Rational Design – Precise, knowledge-driven modifications based on structural insights.
Semi-Rational Design – A blend of targeted changes and evolutionary approaches.
How does computational modeling aid protein engineering?
Computational tools like Rosetta, FoldX, and MOE predict protein structure, stability, and interactions, guiding precise modifications. These tools accelerate protein design by reducing experimental trial-and-error.
What is directed evolution, and why is it important?
Directed evolution mimics natural selection in the lab, generating improved proteins through iterative mutation and screening. Techniques like phage display, yeast surface display, and compartmentalized self-replication (CSR) are revolutionizing drug discovery and enzyme engineering.
How will AI shape the future of protein engineering?
AI-driven tools, such as AlphaFold and machine learning-based design algorithms, are transforming protein engineering by predicting structures with unprecedented accuracy, accelerating discovery, and unlocking novel biomolecular functions.






